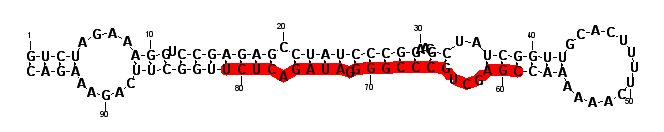| seed | -----------------------------------------------------------------------------------CACCGGG--------------------------------------------------- | 7 | miR-35/36/38/39/40/41 |
| | | | AF16 |
| | ----------------------------------------------------------------------------------TCACCGGGTGAAAA-CTGGTAG------------------------------------- | 21 | 5 |
| | ----------------------------------------------------------------------------------TCACCGGGTGAAAA-CTGGTAGAG----------------------------------- | 23 | 2 |
| cbriggsae | -------------CCGAAGGTCTGGG-----TCGAGCCTCCGCTGGTTTTTTACCGCGGTGATATTCCAAATAAAATGGTTATCACCGGGTGAAAA-CTGGTAGAGG--------TCCGATCCAGTCGTACCATCG----- | |
| cremanei | ------------------TGGTCGGA-----TCGGCCATTCACTGG-TTCTCGCCGCGGTCATATTTCAAAT---ATGATTATCACCGGGTGAAAA-TCAGAGCAGG--------GCCGATCCCACCG------------- | |
| celegans | TTCCCAACTATTATTCTCGGATCAGA-----TCGAGCCATTGCTGG-TTTCTTCCACAGTGGTACTTTCCAT--TAGAACTATCACCGGGTGGAAA-CTAGCAGTGG--------CTCGATCTTTTCC------------- | |
| cbrenneri | --------------TTCAGGGGTGGA-----TCGGGCCTGTGCTGA-TTTTTGCCGCTGTGATATTTCTAAG-CCATGATTATCACCGGGCGAAAATTTGGCACAGG--------CCTCGTCCGCCCTTCCTGAG------ | |
| cjaponica | ----TGCAGTGGTCCCTCGGGGCAGAAGAGCCAGGACCTACATCAGTTTCTTTCCGCGGTAATAAATAATAC---ATTATTATCACCGGGAGAAAAACTGGAGTAGGACCTGCGACTCGACTCGACCTTTATTCCGCTGTA | |
| ppacificus | ------------------GGGATGCA--------------------TATTCTAACCCGGTGACTCAATAAAC--AATGA--GTCACCGGG-------TTAGAATATG--------TCCA---------------------- | |
| | * * * * ** * * ******** * * | |
| cbriggsae | .(((.(((((((( (((..((((..((((.((((((((.((((((((..(((.......))).)))))))))))))))) ))))..)))) ..))))))))....))).))) | 1.000 -57.20 |
| cremanei | .(((.((( ((((((.....(((( ((.(((((.((((.(((..(((... .))).))).))))))))).)) )))).....) )))))))).))). | 1.000 -46.80 |
| celegans | ..................(((..((( ((((((((((((((( ((((..((...((((((.(((.... ..))).))))))..))..)))) )))))))))) ))))))))..))) | 0.989 -54.00 |
| cbrenneri | .((((((((((( (.((((((((((((( ((((((((...((((((..((.... ....)).))))))..)))))))))).)))))))) ))).))))))))...)))). | 1.000 -59.20 |
| cjaponica | ((((((((......((((..((.(((.((((.(((((..((((((.(((((.((((.((.(((((... .))))))).))))))))).))))))..))))).))))...)))...))..))))....)))))))) | 0.964 -52.50 |
| ppacificus | .((..((( ((((((((((((((((((((...... ..))) ))))))))) )))))))))) ))). | 1.000 -42.10 |
| cbriggsae | chromosome:chrX:15336203:15336311:1 | {SimpF: trf} |
| cremanei | chromosome:chrUn:26788903:26788994:-1 | intergenic |
| celegans | chromosome:chrII:11537510:11537620:1 | Opposite_strand|Intronic_coding|Y62F5A.9 ## {MIR: cel-mir-35} |
| cbrenneri | chromosome:chrUn:47821549:47821654:-1 | intergenic ## {SimpF: trf} |
| cjaponica | chromosome:chrUn:117591701:117591834:-1 | Opposite_strand|Intronic_coding|NM_171042 |
| ppacificus | chromosome:chrUn:47519693:47519841:1 | Same_strand|Intronic_coding|NM_001003336 |
| seed | --------------------------------------------------------------------------------CACCGGG--------------------------------------------------- | 7 | miR-35/36/38/39/40/41 |
| | | | AF16 |
| | -------------------------------------------------------------------------------TCACCGGGTG-AAAACTGGTAG------------------------------------- | 21 | 5 |
| | -------------------------------------------------------------------------------TCACCGGGTG-AAAACTGGTAGAG----------------------------------- | 23 | 2 |
| cbriggsae | ----------------AATGGTCTGGGTCGGACCTCCGCTGTTTTTCTCCG-CGGTGATACTCCAAA-GAAAGTCGTTATCACCGGGTG-AAAACTGGTAGAGGTCCG---ATTCAGTCGGACCAT------------ | |
| cremanei | -TCCCTAAT----------------GACGGGAACCGTGCTGGTTTCGAACT-CGGTAATAGACCAGAAGAAAG---CTATCACCGGGTG-GAAACTGGTACGGGTCCCCGAGTCGTGCTATTCTGGA----------- | |
| celegans | -TTCCCAACTATTATTCTCGGATCAGATCGAGCCATTGCTGGTTTCTTCCA-CAGTGGTACTTTCCATTAGAA---CTATCACCGGGTG-GAAACTAGCAGTGGCTCG-------ATCTTTTCC-------------- | |
| cbrenneri | ---CCTGA-----------------GACGGGAACCGAGCTGGTTTCTCTAC-TGGTGATATATCCAATCATAGTGTTTATCACCGGGAG-TAAATTGGCTCGGGTCCC-----------GTCTCGGG----------- | |
| cjaponica | TGCAGTGGTCCCTCGGGGCAGAAGAGCCAGGACCTACATCAGTTTCTTTCCGCGGTAATAAATAATA-CATTA---TTATCACCGGGAGAAAAACTGGAGTAGGACCTGCGACTCGACTCGACCTTTATTCCGCTGTA | |
| ppacificus | ----------------------------GGGATGCATA-----TTCTAACC-CGGTGACTCAATAAACAATGA-----GTCACCGGGT-------TAGAATATGTCCA------------------------------ | |
| | * ** ** * * ******** * * * * | |
| cbriggsae | .(((((((((((((((((((..((..(((((.((. ((((((((....... .........)))))))))).) ))))..))..)))))))) ))....))))))))) | 1.000 -51.49 |
| cremanei | ......(( (((((((.((((((..(((((..((( ((((.((((............ )))).))))))). )))))..)))))).))))...)))))........... | 0.928 -44.10 |
| celegans | ..................(((..((((((((((((((((((((((..((. ..((((((.(((......))) .))))))..)).. )))))))))))))))))) ))))..))) | 0.989 -54.00 |
| cbrenneri | ((((( (((((((.((((((..(((((((..( ((((((((.................)))))))))))) .))))..)))))).)))) )))))))) | 1.000 -56.73 |
| cjaponica | ((((((((......((((..((.(((.((((.(((((..((((((.(((((.((((.((.(((((.. ..))) )))).))))))))).))))))..))))).))))...)))...))..))))....)))))))) | 0.964 -52.50 |
| ppacificus | .((..((((( (((((((( ((((((((((........))) )))))))))) )))))))))))). | 1.000 -42.10 |
| cbriggsae | chromosome:chrX:15336370:15336473:1 | {Repeats: no} |
| cremanei | chromosome:chrUn:26788204:26788308:-1 | intergenic ## {SimpF: trf} |
| celegans | chromosome:chrII:11537510:11537620:1 | Opposite_strand|Intronic_coding|Y62F5A.9 ## {MIR: cel-mir-35} |
| cbrenneri | chromosome:chrUn:47817083:47817176:-1 | intergenic |
| cjaponica | chromosome:chrUn:117591701:117591834:-1 | Opposite_strand|Intronic_coding|NM_171042 |
| ppacificus | chromosome:chrUn:47519696:47519839:1 | Same_strand|Intronic_coding|NM_001003336 |
| seed | ---------------------------------------------------------------------------CACCGGG--------------------------------------------- | 7 | miR-35/36/38/39/40/41 |
| | | | AF16 |
| | --------------------------------------------------------------------------TCACCGGGTG----AAAACTGGTAG---------------------------- | 21 | 5 |
| | --------------------------------------------------------------------------TCACCGGGTG----AAAACTGGTAGAG-------------------------- | 23 | 2 |
| cbriggsae | ------------------CTG--GGTCGGACCTCTGCTGGTATTCTCTGCGATGATAT-TTTGAAAAAAGCTTATCACCGGGTG----AAAACTGGTAGAGGTCCGATCCAGT-------------- | |
| cremanei | ---------------TCCCTAATGACGGGAACCGTGCTGGTTTCGAACTCGGTAATAG-ACCAGAAGAAAGCTATCACCGGGTG----GAAACTGGTACGGGTCC--CCGAGTCGTGCTATTCTGGA | |
| celegans | ---------------GGATCA--GATCGAGCCATTGCTGGTTTCTTCCACAGTGGTAC-TTTCCATTAGAACTATCACCGGGTG----GAAACTAGCAGTGGCTCGATCTTTTCC------------ | |
| cbrenneri | ----------------GGCCG--GATCGGACCTATGCTGGTTTTCACCGCGGTTATAC-TTCTAACAATAATTATCACCGGGTT----AAAACTTGCAGAGGTCCGGTCCGGCC------------- | |
| cjaponica | GATATTCTTGCTCTTTCTCCA--GCTCGGACCTCCGCTGGTTT-CTTTTCGTTGATACGATTTTAATCATGTTATCACCGGGTT----GAAACTGGTAGAGGTCCGACGCTGGAGGCCTCTCATATC | |
| ppacificus | --------TCTGAGTCCCTTC--GATCG--CCTCTGAT------CACTACGATGAAAC-TGAGATAACGAGTTTTCACCGGGTGAACAGAAGCTGTTGAAGAT---ATCGGA--------------- | |
| | * * * * * * * * * ********* ** ** * | |
| cbriggsae | ((( ((((((((((((((..((.(((.((.((.(((((( (((....))))..))))).)))).) )).))..))))))))))))))))). | 1.000 -49.30 |
| cremanei | ......(((((((((.((((((..(((((..(((((((.(((( ............)))).))))))). )))))..)))))).))) )...)))))........... | 0.928 -44.10 |
| celegans | (((..( (((((((((((((((((((((..((...((((((. (((......))).))))))..)).. ))))))))))))))))))))))..))) | 0.992 -53.70 |
| cbrenneri | ((((( ((((((((((.(((.((((((.(((.((((.(((. ((........)).))).))))))). )))))).))).))))))))))))))) | 1.000 -51.90 |
| cjaponica | ...............(((((( (((((((((((..((((((( (..((((.(((((.(((....)))....))))).)))).. ))))))))..))))))))).))))))))........... | 0.974 -52.40 |
| ppacificus | (((((....(((( (((.( (.((((.( ((((.((.(((((( (.........))))).)).))))))).)))).)).))))))).. .))))) | 1.000 -30.60 |
| cbriggsae | chromosome:chrX:15336518:15336605:1 | {Repeats: no} |
| cremanei | chromosome:chrUn:26788204:26788308:-1 | intergenic ## {SimpF: trf} |
| celegans | chromosome:chrII:11537528:11537620:1 | Opposite_strand|Intronic_coding|Y62F5A.9 ## {MIR: cel-mir-35} |
| cbrenneri | chromosome:chrUn:47823952:47824042:-1 | intergenic |
| cjaponica | chromosome:chrUn:117592041:117592160:-1 | Opposite_strand|Intronic_coding|NM_171042 |
| ppacificus | chromosome:chrUn:171595123:171595212:-1 | intergenic ## {Repeats: (GA)n 2 38 -1 class=Simple_repeat} ## {SimpF: trf} |
| seed | -------------------------------------------------------------AGCUGCC------------------------------ | 2 | novel |
| seed | ------------------------------------------------------------GAGCUGC------------------------------- | 1 | novel |
| | | | AF16 |
| | ------------------------------------------------------------GAGCTGCCCGGGGATAGACTCT---------------- | 22 | 2 |
| | -----------------------------------------------------------CGAGCTGCCCGGGGATAGACTCT---------------- | 23 | 1 |
| cbriggsae | GTCTAGAAAGGT-CCGAGAGCCTATCCCGGAACGCTATCGGTTGCACTTTTCAAAAAACCGAGCTGCCCGGGGATAGACTCTTGGC-TTCAGAAAGAC | |
| cremanei | ----------------AGAGTTTATCCAGGAAAGCTATCGGCCGAGTACTT---AGAACCGAGCTGCCCTCGGATAGACTCT---------------- | |
| celegans | TTCTTGAAAACT-CCAATAGTTTCTCTGGGAAAGCTATCGGCCAAA---TTTAACTGTCCGAGCTGCCCTCAGAAAAACTCTTGGC--TCATCGAGAA | |
| cbrenneri | GTCTCGAACACCACCAAGAGTTTATCCAGGAAAGCTATCGGACTGACATTT---AGTACCGAGCTGCCCTGGGATAGACTCTTGGTAATTTTAGAGAC | |
| | * ** * ** **** ******** ** *********** ** * ***** | |
| | +++++++++++++++++ +++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++ ++++++++++++++++ | NM_001047975 |
| | +++++++++++++++++ +++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++ ++++++++++++++++ | NM_001047939 |
| | +++++++++++++++++ +++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++ ++++++++++++++++ | NM_101724 |
| | ----------------- ------------------------------------------------------------------------- ---------------- | NM_001011629 |
| | +++++++++++++++++ +++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++ ++++++++++++++++ | NM_001058128 |
| | +++++++++++++++++ +++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++ ++++++++++++++++ | NM_063457 |
| | +++++++++++++++++ +++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++ ++++++++++++++++ | NM_068373 |
| | +++++++++++++++++ +++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++ ++++++++++++++++ | NM_067556 |
| | +++++++++++++++++ +++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++ ++++++++++++++++ | NM_202130 |
| | ----------------- ------------------------------------------------------------------------- ---------------- | NM_014971 |
| | +++++++++++++++++ +++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++ ++++++++++++++++ | NM_062799 |
| cbriggsae | ((((....(((. (((((((.(((((((((...((..((((((.............))))))...))))))).)))).)))))))) )).....)))) | 1.000 -37.12 |
| cremanei | ((((((((((((((..(((..((((.......... ....)))))))..))).))))))))))) | 1.000 -25.74 |
| celegans | .((((((..... ((((.(((((.((((((..(((..((((.((.. ......)).)))))))..)).)))).))))).)))). ...)))))). | 1.000 -31.00 |
| cbrenneri | (((((.......((((((((((((((((((..(((..(((((((((...)) ))).)))))))..))))).))))))))))))).......))))) | 1.000 -46.14 |
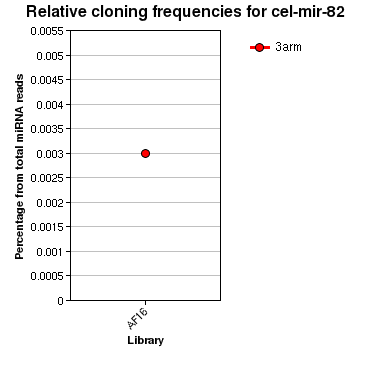
| seed | ----------------------------------------------------------------------GAGAUCA------------------------------------------------ | 1 | miR-80/81/82 |
| | | | AF16 |
| | ---------------------------------------------------------------------TGAGATCA------------------TTCAAGAAGCCC------------------ | 20 | 1 |
| cbriggsae | -------------TGAAATTTTCAATACGTTGGCTTTTTGGCTGGTCTCATACCGTTTAGTACTGGATATGAGATCA------------------TTCAAGAAGCCCAAAAAAGAGTTCATTTCA | |
| cremanei | -----------CTTAGGTCATGTCTTTCACTCACTTTTTG----------------------------ATGAGATCA------------------TTCATGCACCTGCC---------------- | |
| celegans | TGAACTCATGAAATAGGTTCTTTTAGCAACCGGTTTTCTCTGTGATCTACAGAATGACAGCTAATCGTCTGAGATCATCGTGAAAGCCAGTTGTTTTTATGAACTCTTATCAAGAA---AATTCA | |
| cbrenneri | -----CCGGGTTGTAGAATTCTTCAGTAATTAGTATCTTGTACA-------------------TAACTGTGAGATCA------------------TTCAACAGCCTTC----------------- | |
| ppacificus | -----------------CTCTCAATTTCGCCTGCTCTCTCGTCA------------TCTCTGATCGATGAGAGATCA------------------TTCGAAA-------TGAAGAG--------- | |
| | * ******* ** | |
| | +++++ ++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++ +++++++++++++++++++++++++++++++++++ | NM_071218 |
| | +++++ ++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++ +++++++++++++++++++++++++++++++++++ | NM_067556 |
| | +++++ ++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++ +++++++++++++++++++++++++++++++++++ | NM_001083250 |
| | ----- ---------------------------------------------------------------- ----------------------------------- | NM_001085890 |
| | +++++ ++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++ +++++++++++++++++++++++++++++++++++ | NM_070819 |
| | +++++ ++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++ +++++++++++++++++++++++++++++++++++ | NM_002710 |
| cbriggsae | .(((((.(((........(((((((((..(((((((((((((........))).)))))))))) ..))))))))).......)))...))))). | 0.980 -31.86 |
| cremanei | ..(((((((((..(((((.(((.....)) ))))))... ..)))).))))).. | 1.000 -11.80 |
| celegans | .(((.((.(((...((((((....((((((.(((((((...(((((((.(((.((((.......))))))))))))))...))))))).)))))).....)))).))..))).)). ..))). | 0.989 -37.40 |
| cbrenneri | ..(((((((.((((((((((((.((..((((...))))) ).)))).))))..) ))).))))))).. | 1.000 -16.90 |
| ppacificus | ((((..((((((..((.(((((((((( ((...))).))))))))).)) ..))))) )..)))) | 1.000 -25.10 |
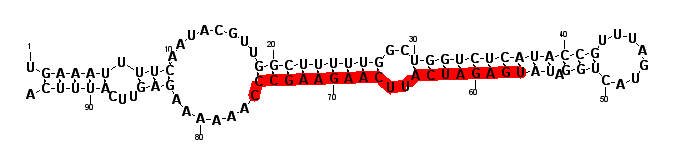
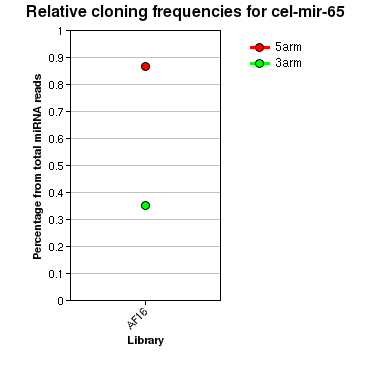
| seed | --------------------------------AUGACA------C-------------------------------------------------------------------------------------------------- | 332 | miR-64 |
| seed | ---------------------------------------------------------------------------------CUACCGG------------------------------------------------------- | 121 | novel |
| seed | -------------------------------------------------------------------------------------CGGGUUC--------------------------------------------------- | 8 | novel |
| seed | ----------------------------------------------------------------------------------UACCGGG------------------------------------------------------ | 6 | novel |
| | | | AF16 |
| | -------------------------------CATGACA------CTGAAGCGGGAATGG------------------------------------------------------------------------------------ | 22 | 310 |
| | --------------------------------------------------------------------------------CCTACCGGGTTCAGTGTCATACT---------------------------------------- | 23 | 114 |
| | -------------------------------CATGACA------CTGAAGCGGGAATG------------------------------------------------------------------------------------- | 21 | 9 |
| | -------------------------------CATGACA------CTGAAGCGGGAAT-------------------------------------------------------------------------------------- | 20 | 9 |
| | ------------------------------------------------------------------------------------CCGGGTTCAGTGTCATACT---------------------------------------- | 19 | 7 |
| | ---------------------------------------------------------------------------------CTACCGGGTTCAGTGTCATACT---------------------------------------- | 22 | 6 |
| | --------------------------------------------------------------------------------CCTACCGGGTTCAGTGTCATACTT--------------------------------------- | 24 | 4 |
| | -------------------------------CATGACA------CTGAAGCGGGAA--------------------------------------------------------------------------------------- | 19 | 3 |
| | --------------------------------------------------------------------------------CCTACCGGGTTCAGTGTCAT------------------------------------------- | 20 | 2 |
| | -------------------------------CATGACA------CTGAAGCGGGAATGGAA---------------------------------------------------------------------------------- | 24 | 1 |
| | --------------------------------------------------------------------------------CCTACCGGGTTCAGTGTCATAC----------------------------------------- | 22 | 1 |
| | ------------------------------------------------------------------------------------CCGGGTTCAGTGTCATACTT--------------------------------------- | 20 | 1 |
| cbriggsae | -----------------AGACCCTCACCAATCATGACA------CTGAAGCGGGAATGGAAGCGTAGTCAGATTGCAATTCCTACCGGGTTCAGTGTCATACTTGGTGCG---------------GATGAGTCTG-------- | |
| cremanei | -------------------------------CATGACA------CCGATCTCGCGATGG---------------TCGTTTTCCAAAGCGATCGATAAGGCGGTAG--------------------AGTTGTCCTG-------- | |
| celegans | GTGCTCTTGGAGCCATGGAGCCTTCGCCGATTATGACA------CTGAAGCGTAACCGAACACCATAT----TTTGAGATTCTGCTACGCGCAGTGCCATGCTCGGCGCGTTGGCTCCATTAAAAAACGGCCACAAAAATGCC | |
| cbrenneri | -------------------------------AATGACA------CTGAAGTGGG--------------------CCAAGTTTTGAGTTTTTTTTTGGAACTTTG---------------------AGTGCTCATA-------- | |
| cjaponica | -------------------------------CATGACA------CTGACAATAAGGAGG-------------TTGGGAGCTCTACGTACCTAAACTATATT------------------------AAAGTGCATA-------- | |
| ppacificus | -------------------------------CATGACACATCGCCTGAGAATCAATCGAA--------------TTAAGTTTCATTTCGCTAATTGAATCTCAATT-------------------GAGTTTCATA-------- | |
| | ****** * ** | |
| | +++++ +++++++++++++++++++++ ++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++ ++++++++++ +++++ | NM_001102902 |
| | ----- --------------------- ------------------------------------------------------------------ ---------- ----- | NM_001077612 |
| | +++++ +++++++++++++++++++++ ++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++ ++++++++++ +++++ | NM_065190 |
| | ----- --------------------- ------------------------------------------------------------------ ---------- ----- | NM_001129219 |
| cbriggsae | ((((((.((((((..(((((( (((((.(((....((((..(((((...)))))..))))..)))..)))))))))))..)))))).) )....)))). | 1.000 -40.60 |
| cremanei | ((.(((( ....((((((.(((( ((((((....)))))))).))...)))..)) )..)))).)) | 0.964 -14.90 |
| celegans | ..((..(((..(((((((((((..((((((.((((.(( (((..(((((...(((...((... ..))...)))...))))).))))).)))).))))))....))))))))........)))..)))....)). | 0.957 -49.40 |
| cbrenneri | .(((((( (((((((... (((((.............))))).))))). )))).)))). | 0.997 -14.62 |
| cjaponica | .(((.(( ((...((((.((... (((((.((.....)).))))).)).)))) ..))))))). | 1.000 -13.00 |
| ppacificus | .((((.((.(((..(((((.((((((((( ...........))))...))))).)))))..) )))).)))). | 0.999 -16.70 |
| cbriggsae | chromosome:chrIII:218506:218602:-1 | Same_strand|Intronic_non-coding|NM_001102902|LOC100124966 ## Opposite_strand|Intronic_coding|NM_001077612|zgc:153009 ## Opposite_strand|Intronic_non-coding|NM_001129219|nhr-121 ## {Repeats: no} |
| cremanei | chromosome:chrUn:30737923:30738059:1 | Opposite_strand|Boundary_non-coding|NM_181404 ## Opposite_strand|Boundary_coding|NM_061875 ## Same_strand|Intronic_coding|NM_182266 |
| celegans | chromosome:chrIII:2172976:2173108:1 | intergenic ## {MIR: cel-mir-65} |
| cbrenneri | chromosome:chrUn:96785595:96785731:-1 | intergenic |
| cjaponica | chromosome:chrUn:125505645:125505781:-1 | Same_strand|Intronic_coding|NM_067721 |
| ppacificus | chromosome:chrUn:116309862:116309998:-1 | Opposite_strand|Intronic_coding|NM_031126 ## Opposite_strand|Boundary_coding|NM_001029437 ## Same_strand|Intronic_coding|NM_198493 |